Introduction
When you combine datasets, run downstream analyses, or visualize results, you almost always need your genes in a single, consistent identifier system (e.g., Ensembl IDs or Entrez IDs). Unfortunately, public resources and pipelines use many different “names” for the same gene: Ensembl stable IDs, Entrez Gene IDs, HGNC symbols, UniProt accessions, microarray probe IDs, etc.
g:Profiler (g:Convert) solves this by providing a central, regularly updated service that translates gene identifiers across dozens of namespaces for hundreds of organisms. The task below wraps g:Convert to give you a simple, reproducible way to normalize your IDs from a table.
🧰 Prerequisites
- Access to Constellab and a valid Digital Lab environment
- Installed bricks: gws_omix ≥ 0.13.10
- Differentially expressed genes file (csv or tsv) or Table , a list of gene symbols, Ensembl IDs, Entrez IDs, UniProt accessions
🧪 Workflow: Step by Step
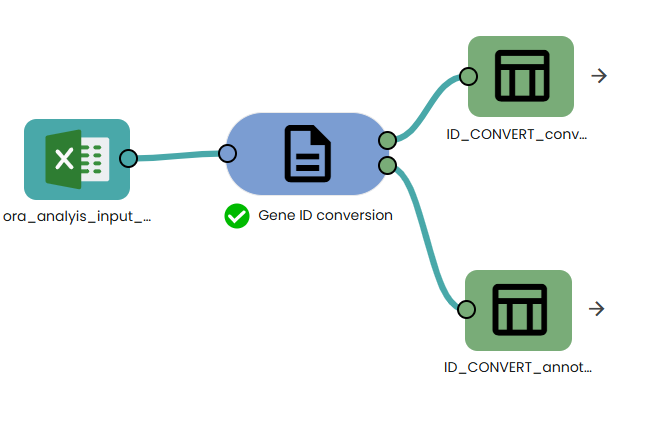
- Add the required task : Gene_ID_conversion
This task performs identifier harmonization on a user-provided list of gene IDs. It uses the g:Profiler g:Convert service to translate input identifiers (e.g., gene symbols, Ensembl IDs, Entrez IDs, UniProt accessions) into a selected target namespace. Unlike manual mapping or static resources, g:Convert queries a continuously updated central database, ensuring that results reflect the latest stable identifiers and gene annotations.
2. Configurer the task: please read carefully the documentation
3. run task
Information :
Parameters
Outputs

