Introduction
Contextualizing a metabolic network involves understanding its relevance within a broader biological system or specific research context. By contextualizing a metabolic network, researchers can gain insights into how it operates within living organisms, its role in various cellular processes, and its implications for health and disease.
In Constellab you can contextualise your metabolic network using a context. To create a context you can use the Task Context Builder, which we will see in more detail in this documentation.
Data upload
First you need to upload a metabolic network. In Constellab you can contextualise the reactions and/or metabolites.
Contextualise the reactions
To contextualise the reactions, you need to create a Flux table importer. To do this you need to import a File like this:
reaction_id,target,lower_bound,upper_bound,confidence_score
EX_o2_e,,0,1000,1
Contextualise the metabolites
To contextualise the metabolites, you need to create a Phenotype table importer. To do this you need to import a File like this:
id,target,lower_bound,upper_bound,confidence_score,chebi_id
glc__D_e,,0,1000,1.0,CHEBI:12965
o2_c,,0,1000,1.0,CHEBI:7860
etoh_c,1.23,0,2.0,1.0,CHEBI:42377
Multiple simulations
These Tasks manages multiple simulations. So if you have different values of target,lower_bound,upper_bound; set them as a list like this:
id,target,lower_bound,upper_bound,confidence_score
id,"[None, 0.045, 0.035]","[0.01, 0.008, -0.02]","[0.03, -0.003, 0.001]","[1, 1, 1]"
Protocol
First you need to import all the different files. Then you can connect all the resources to the Task Twin Builder.
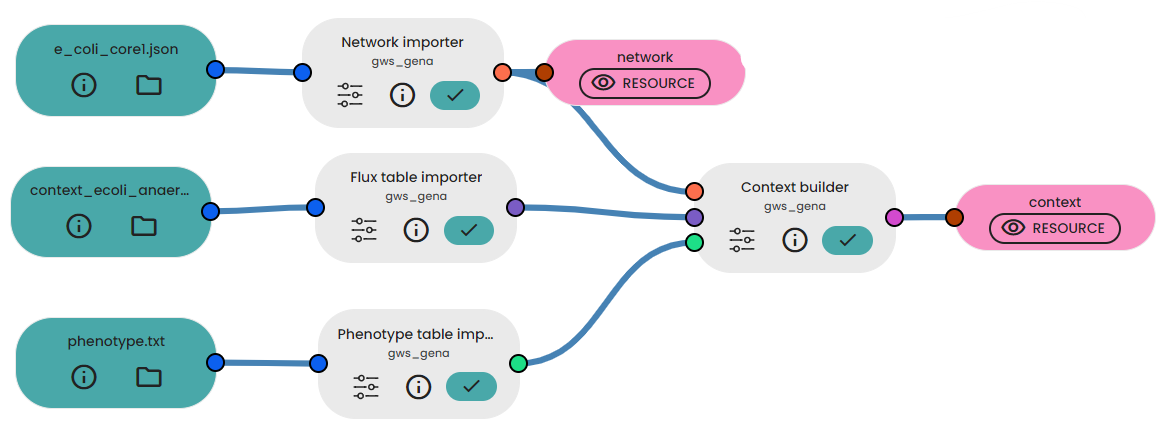
In the output, you will get a context with the measures of reactions fluxes and metabolites, see below a screenshot.

You are now ready to use the context you have created to contextualise your metabolic network using the Twin Builder task. You can then use FBA, FVA tasks, for example, to exploit your twin.

