Introduction
When you open an annotated genome file, most of the information is there, but the structure is still hard to see.
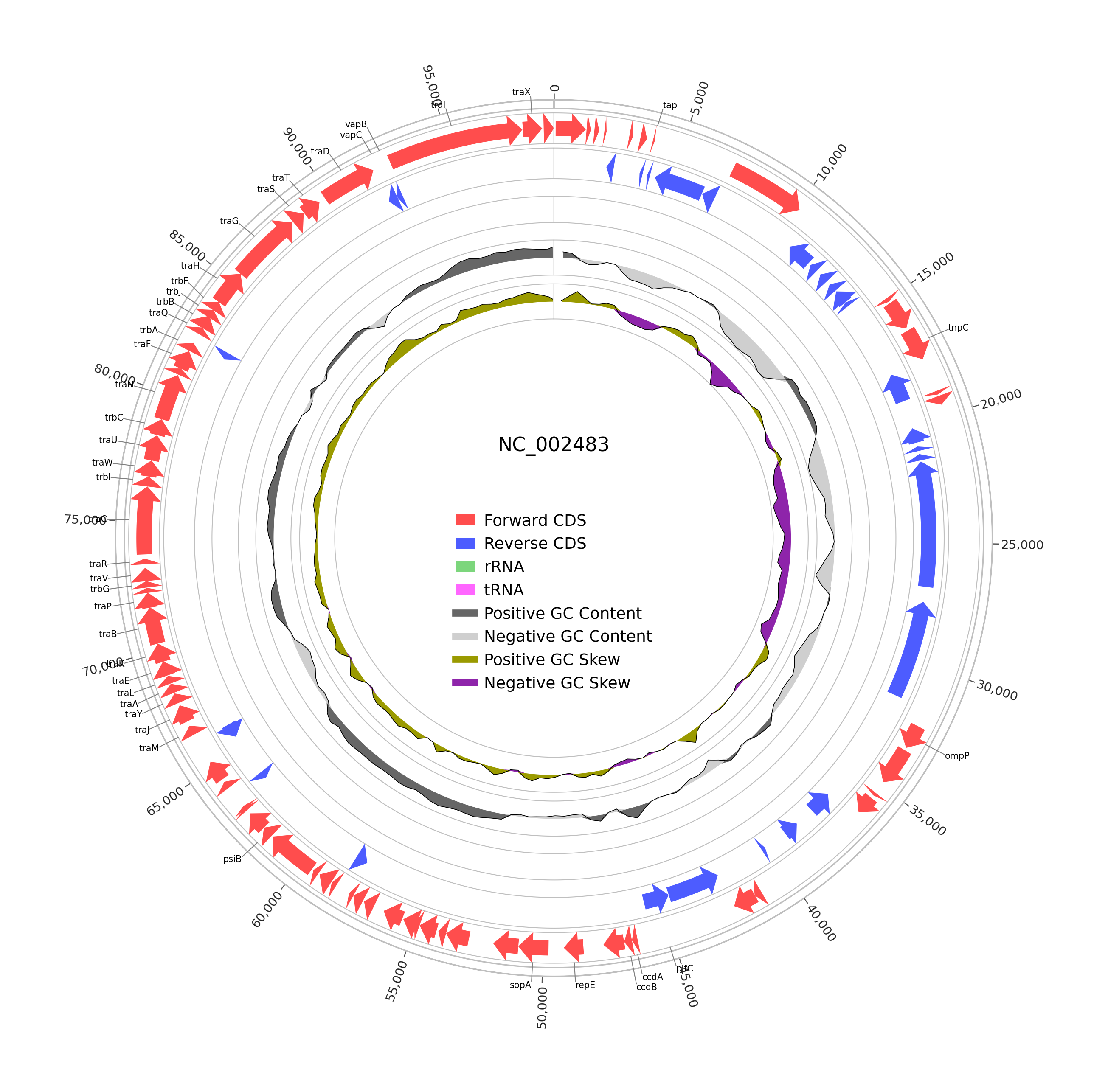
Genes, RNAs, coordinates, and strand orientation are buried in the file format. This task wraps pyCirclize to turn those annotations into clean circular genome plots that are much easier to inspect and share.
In this wrapper, the task is focused on annotated genome circular maps from GenBank or GFF3 files. It automatically scans a file or a folder, keeps supported genome records, and generates one PNG circular plot per input genome file.
🧰 Prerequisites
- Access to Constellab and a valid Digital Lab environment
- Installed bricks: gws_omix ≥ 0.13.12
- A supported annotated genome file, or a folder containing supported genome files
- Supported input formats: GenBank (
.gb, .gbk, .genbank, .gbff) or GFF/GFF3 (.gff, .gff3)
🧪 Workflow: Step by Step
1. Add the required task: Circular Genome Visualization (pyCirclize)
This task performs circular genome visualization on one annotated genome file or on a folder of genome files. It uses pyCirclize to generate a compact circular representation of genome structure and annotations. In this wrapper, the task is described as “Circular Genome Visualization (pyCirclize)” .
2. Configure the task
Provide the genome_input.
The task accepts:
- a single file, or
- a folder containing one or more supported files.
3. Run the task
The task scans the input, loads supported files, extracts annotated features, and generates one circular PNG plot for each successfully processed genome file. Files with no supported annotations are skipped.
Information
This wrapper focuses on common annotation features:
- CDS
- rRNA
- tRNA
- tmRNA
- ncRNA
When usable sequence data is available, the plot can also include:
If a GenBank record has no usable sequence, GC-based tracks are skipped.
If a file contains no supported annotation features, it is skipped and no PNG is produced for that file.
Outputs