Introduction
Quality control and preprocessing of sequencing data are crucial steps in NGS data analysis pipelines, as they help improve the accuracy and reliability of subsequent analyses such as genome assembly, variant calling, and gene expression quantification. Trimmomatic addresses these needs by providing a comprehensive set of algorithms and functions for read trimming and filtering.
Main functions of trimmomatic
Its primary purpose is to remove low-quality bases, adapter sequences, and other artifacts from raw sequencing reads, resulting in cleaner and more reliable data for downstream analysis.
Steps to follow
Ensure that the
- version 0.1.2 brick is loaded
- So first, upload your fastqc folder to the Databox.
- Then, create a new experiment.
- Import your resource
Link it to the
task available in the
- brick.
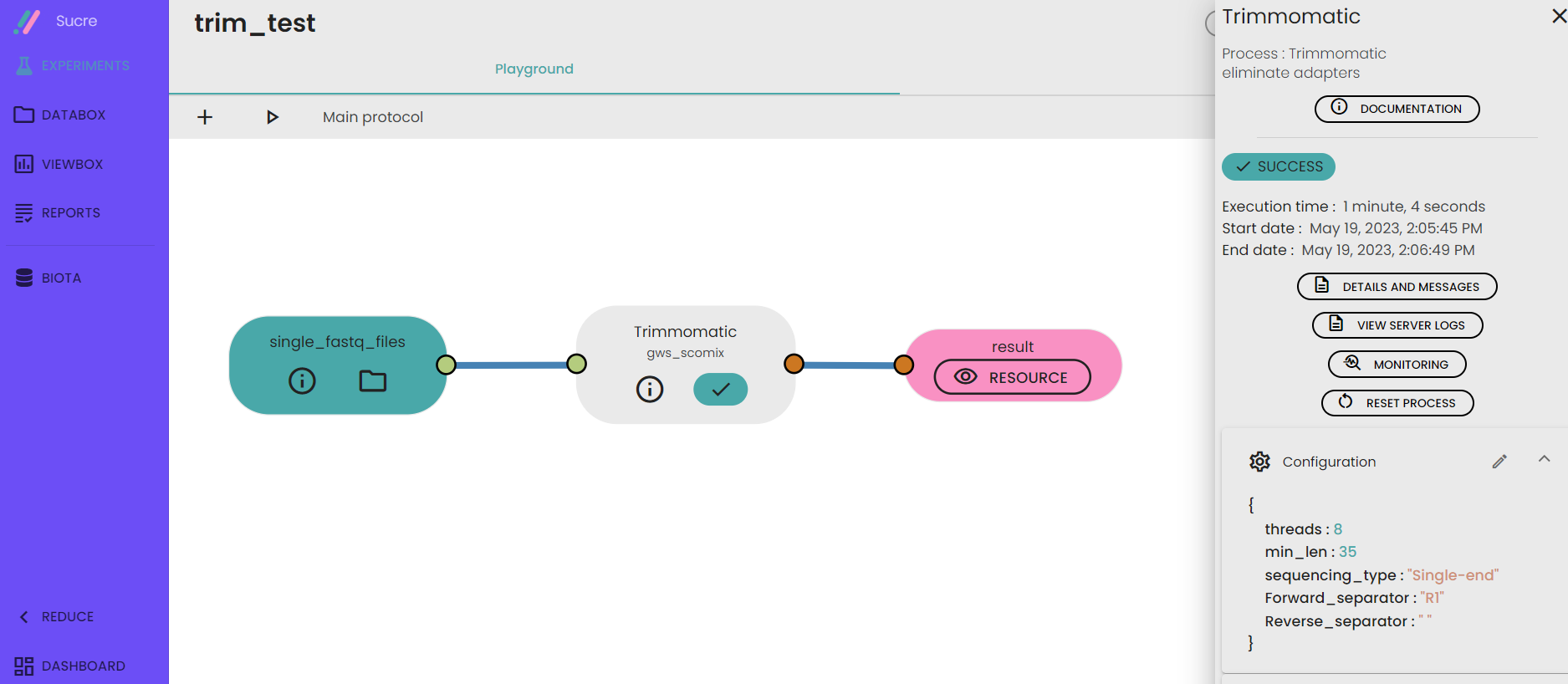
- Specify the type of sequencing "single end" or "paired end" , forward and reverse file differentiator , number of threads and minimum length of reads to be kept.
- Run your experiment
- A folder containing trimmed data wil be generated

Comments (0)
Write a comment